Mapping functional interactions with fMRI
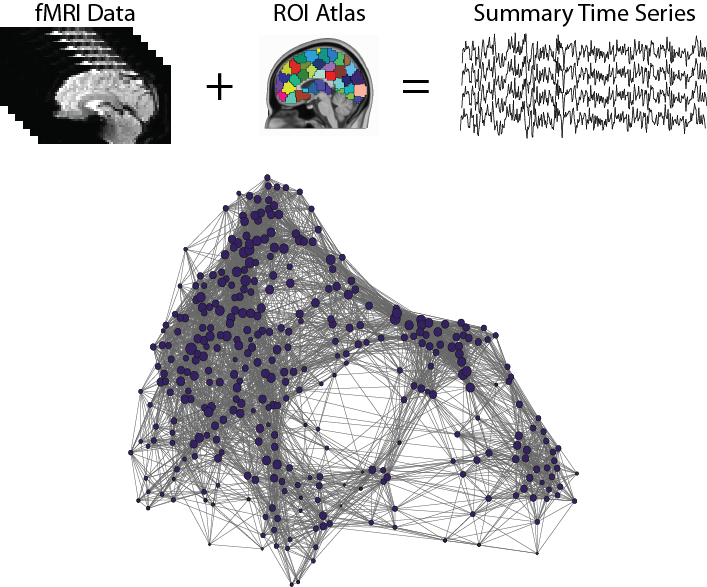
R. Cameron Craddock, PhD
Research Scientist VI, Nathan S. Kline Institute for Psychiatric Research, New York, NY
Director of Imaging, Child Mind Institute, New York, NY
July 29, 2014

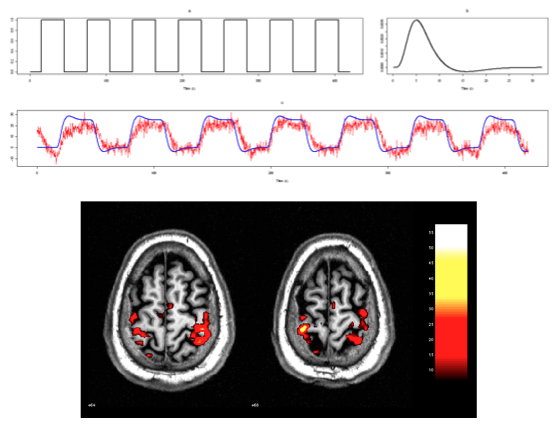
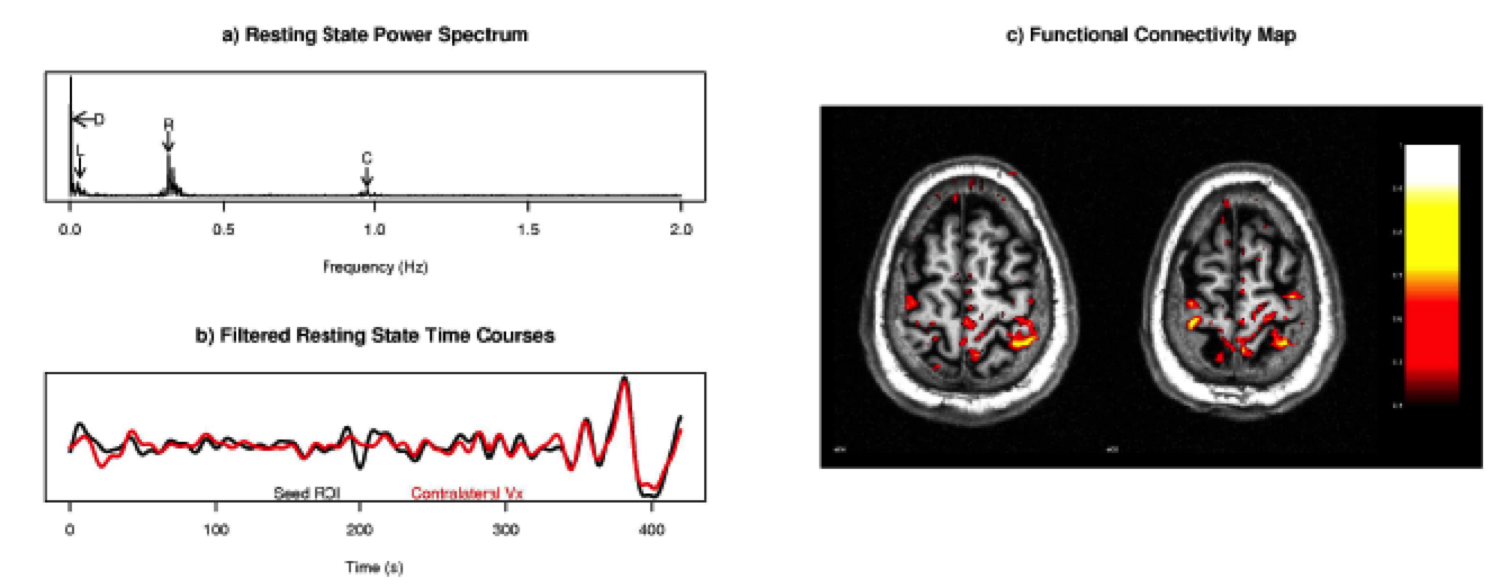
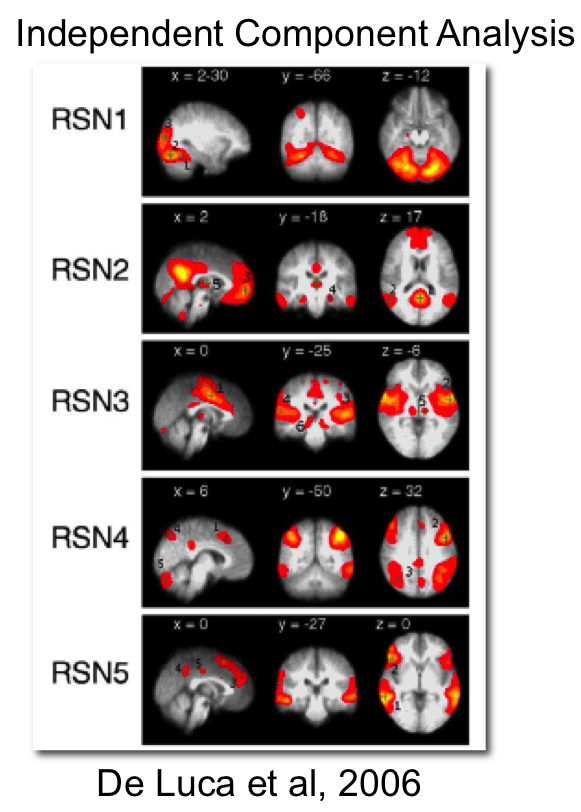
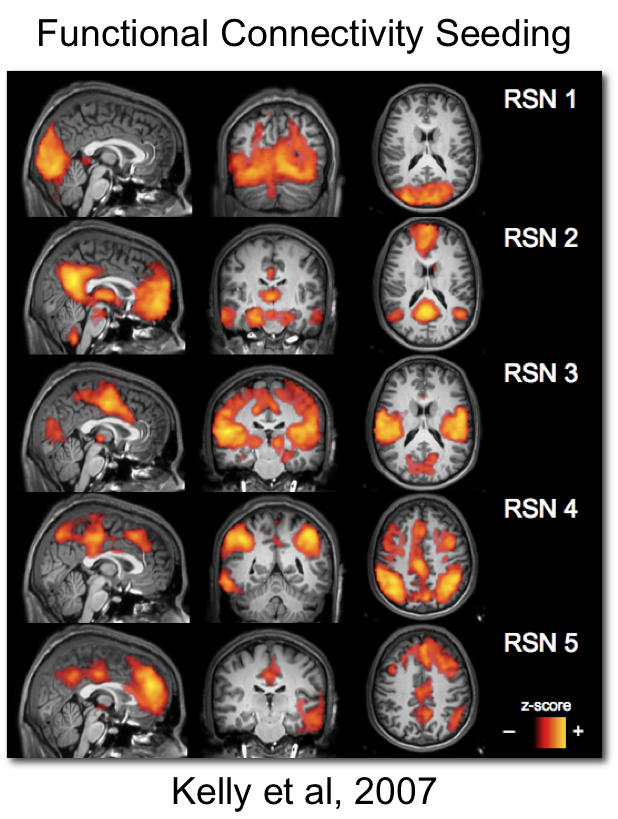
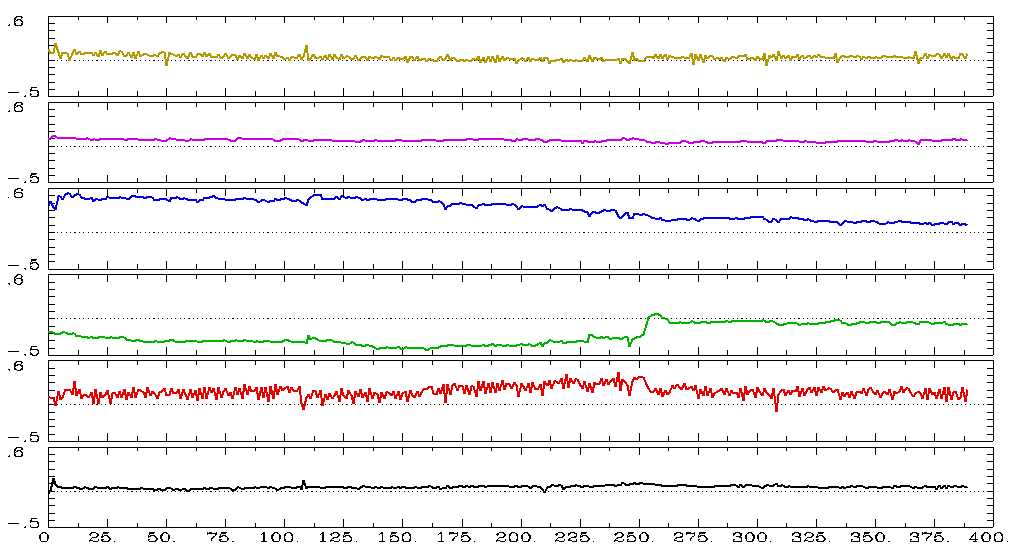
\[v[t] = \mathbf{\beta}^T\mathbf{\eta[t]} + \nu[t] \]